A recurring theme in evolution is how existing components in a biological system can be modified or reused for another context or function. Take, for example, signaling pathways. In complex organisms, such as humans, there are thousands of processes that require coordination via signaling, such as developmental programs, immune processes, homeostatic regulation, etc. Yet, signaling in these organisms has evolved so that remarkably few of the highly conserved signaling pathways govern multitudes of aforementioned processes. Part of the ongoing question in biology involves how a single pathway is able to support such a variety of processes.
One could imagine a number of ways this could be made possible: contexts outside the signaling pathway, such as cell state or epigenetic modification of the genome, could direct signaling to take effect in a specified manner. Another way is through introducing layers of complexity in the signaling pathway itself. Adding complexity to how a signal is transmitted or transduced can broaden the information capacity and enable the signaling to be used in more contexts. For example, signaling systems often have multiple ligands and receptors, some of which are only used for specific purposes or create a unique signaling response. In our study, we discovered how the canonical Wnt pathway has implemented a layer of complexity through the use of two distinct but opposing types of transcriptional elements that respond to a single signal, allowing rich variation of transcriptional output from a monotonic input (Kim, Cho, Hilzinger, et al., 2017).
The canonical Wnt pathway works through a messenger protein called β-catenin, which is in constant flux of synthesize and rapidly degradation in a cell. Signal activation by Wnt proteins inhibits degradation of β-catenin, which allows the protein to accumulate and localize to the nucleus with its binding partner, Tcf. Traditionally, it is understood that the Tcf/β-catenin complex binds to a specific DNA element- aptly called the Wnt-Responsive Element (WRE), recruits transcription factors, and activates gene expression. Barring crosstalk and feedback mechanisms, this signal transduction only allows monotonic response to signaling by Wnt proteins: β-catenin increase leads to activation of a target gene.
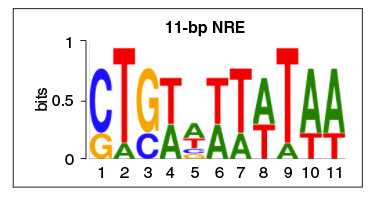
However, we’ve discovered another DNA element that recruits β-catenin/Tcf to suppress gene activation, named 11-bp Negative Regulatory Element (11-bp NRE), in a recently published work that was began with Lea’s groundwork in Marc Kirschner’s lab, and completed with major efforts from Jae Cho, Tom Hilzinger, and several others from our own lab,. We have identified these 11-bp NREs in Wnt target genes from humans to mice to frogs, and demonstrated that they play important roles in regulation of these genes.
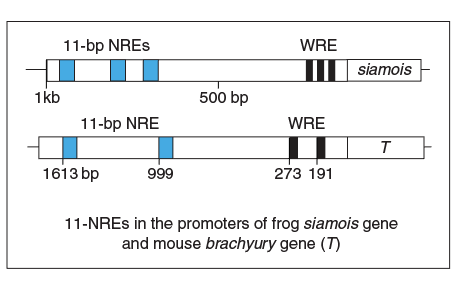
The fascinating part about the discovery of the 11-bp NREs is that they are located closely to the traditional WREs in a gene’s promoter, and work in tandem in regulating gene expression. Here, we have two elements that both recruit β-catenin, one activating a gene and the other suppressing it.
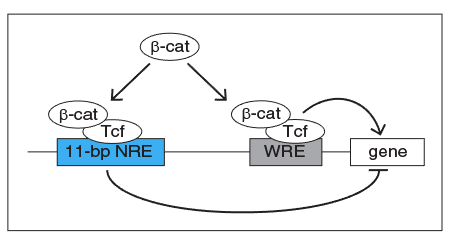
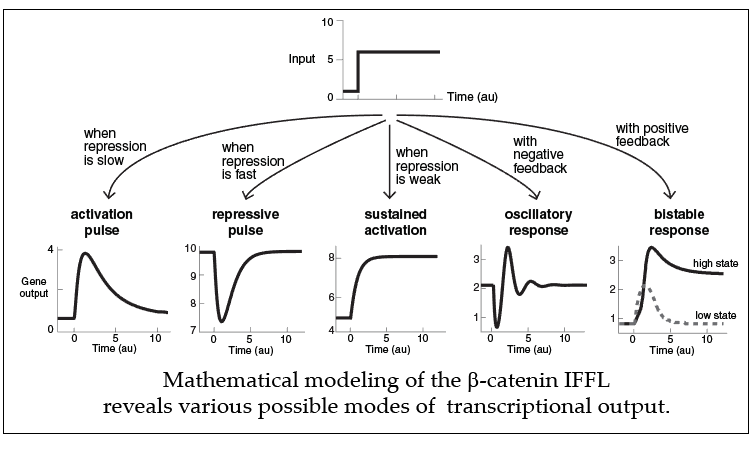
How does this paradoxical interaction work to regulate genes? This type of regulation is actually found frequently and well characterized as a network motif in biology, identified as Incoherent Feedforward Loop (IFFL). Modeling simulations by Harry Nunns in our lab demonstrate that the IFFL composed of the two elements in the Wnt pathway can generate a diverse array of transcriptional responses, such as pulsing, oscillations, bistable response, or fold change detection. Thus, presence of two distinct elements to transduce information by b-catenin allows versatility in how the Wnt signaling pathway controls the response of its target genes, which suggests a layer of mechanism for how the Wnt signaling pathway can function in various contexts to produce a rich variety of dynamic responses.
— Kibeom Kim
Reference: Two element pathway in the Canonical Wnt Pathway. (Current Biology, Volume 27, Issue 15, p2357–2364.e5, 7 August 2017)
Cover photo designed by Andrew Liu